While significant strides have been made in predicting static protein structures, understanding protein dynamics, influenced by ligands, is essential for grasping protein function and advancing drug discovery. Traditional docking methods often treat proteins as rigid, limiting their accuracy. Although molecular dynamics simulations can suggest relevant protein conformations, they are computationally intensive. Recent advances, such as AlphaFold, predict structures from sequences but generate only a few conformations, missing the dynamic nature of proteins. This limitation affects docking accuracy, as AlphaFold-predicted structures may not reflect the optimal configurations for ligand binding, leading to inaccurate predictions.
Researchers from Galixir Technologies, School of Pharmaceutical Science, Sun Yat-sen University, Center for Theoretical Biological Physics and Department of Chemistry, Rice University, and Global Institute of Future Technology, Shanghai Jiao Tong University have developed DynamicBind, a deep learning method that uses equivariant geometric diffusion networks to create a smooth energy landscape, enabling efficient transitions between different equilibrium states. DynamicBind accurately predicts ligand-specific conformations from unbound protein structures without holo-structures or extensive sampling. It excels in docking and virtual screening benchmarks, accommodating large protein conformational changes, and identifying hidden pockets in new protein targets. This method shows promise in accelerating the development of small molecules for previously undruggable targets, advancing computational drug discovery.
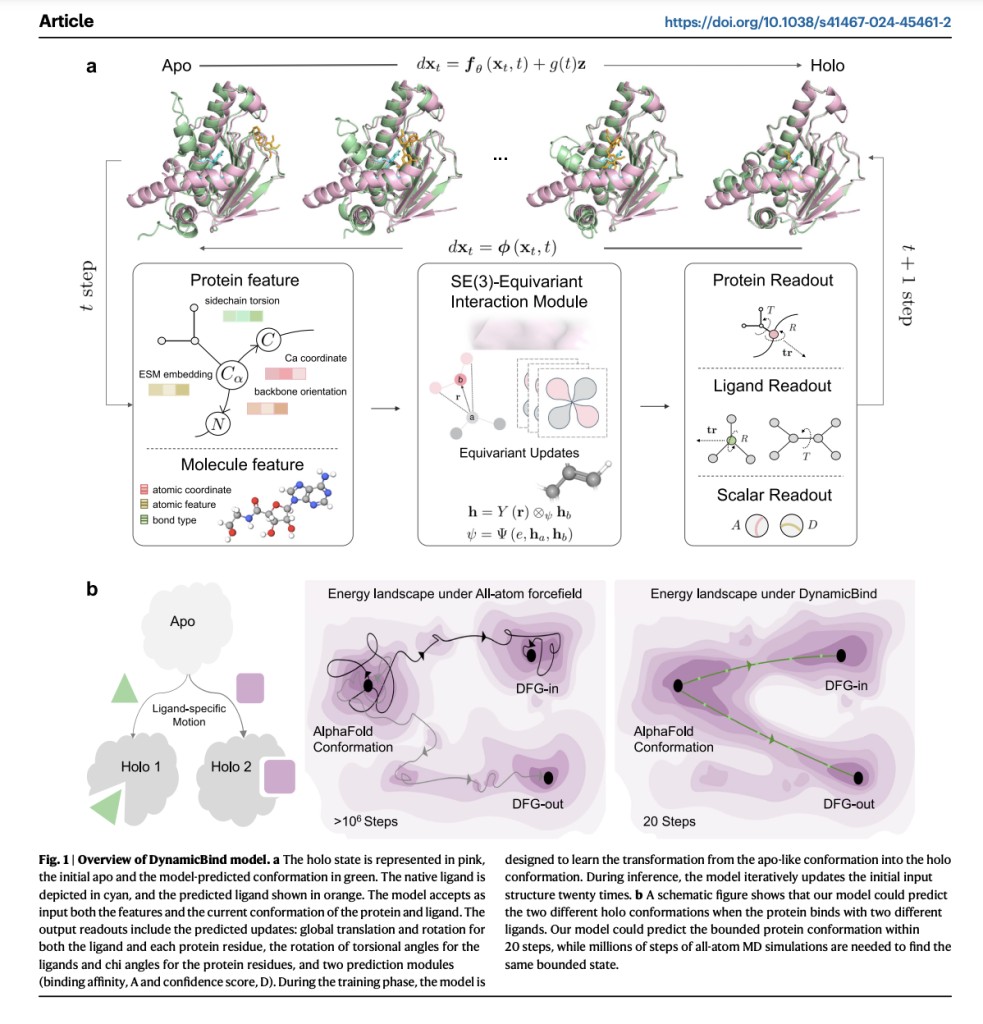
DynamicBind, a geometric deep generative model for dynamic docking, efficiently adjusts protein conformations from initial AlphaFold predictions to holo-like states. It handles significant conformational changes, like the DFG-in to DFG-out transition in kinases, better than traditional molecular dynamics simulations. DynamicBind achieves this by learning a funneled energy landscape that minimizes frustration during transitions between biologically relevant states. Unlike traditional Boltzmann generators, DynamicBind is generalizable to new proteins and ligands.Â
The DynamicBind model is an E(3)-equivariant, diffusion-based graph neural network using a coarse-grained representation to predict protein-ligand binding conformations. It efficiently transforms input structures to account for 3D trans-rotational and parity changes, outperforming traditional methods with less data. The model employs a morph-like transformation for training, interpolating between crystal and AlphaFold structures. Utilizing a graph representation, each protein residue and ligand atom is a node with various features. DynamicBind updates these nodes through tensor products and diffusion processes to predict side-chain dihedrals, torsion angles, translations, and rotations, enhancing binding affinity predictions.
DynamicBind is a versatile tool for predicting protein-ligand complex structures, adept at accommodating significant protein conformational changes. During inference, it gradually adjusts ligand positions and internal angles over 20 iterations while also adapting protein conformations, particularly side-chain angles, improving upon AlphaFold-predicted structures. Unlike traditional models, it employs a morph-like transformation rather than Gaussian noise perturbations, which enhances the model’s ability to capture biologically relevant conformational changes. DynamicBind excels in ligand pose prediction, reducing clashes and revealing cryptic pockets, as demonstrated across various benchmarks and case studies, showcasing its potential for drug discovery applications.
In conclusion, DynamicBind integrates protein conformation generation and ligand pose prediction into a single, end-to-end deep learning framework, significantly faster than traditional MD simulations. Unlike conventional docking methods that require predefined binding pockets, DynamicBind performs global docking, which is ideal for identifying cryptic pockets. This reduces potential side effects by targeting specific proteins and aids drug discovery by predicting unintended protein targets or identifying targets in phenotype screening. Although it shows excellent performance, improvements are needed for better generalization to proteins with low sequence homology. Advances in Cryo-EM and incorporating binding affinity data can enhance DynamicBind’s capabilities.
Check out the Paper and GitHub. All credit for this research goes to the researchers of this project. Also, don’t forget to follow us on Twitter. Join our Telegram Channel, Discord Channel, and LinkedIn Group.
If you like our work, you will love our newsletter..
Don’t Forget to join our 42k+ ML SubReddit
The post DynamicBind: A Deep Learning Approach for Dynamic Protein-Ligand Docking and Drug Discovery appeared first on MarkTechPost.
Source: Read MoreÂ